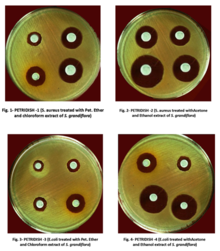
Disk diffusion test
In diagnostic labs, the assay is used to determine the susceptibility of bacteria isolated from a patient's infection to clinically approved antibiotics.
When bioprospecting, the assay can be performed with paired strains of bacteria to achieve dereplication and provisionally identify antibacterial mechanism of action.
[6][7] In diagnostic laboratories, the test is performed by inoculating the surface of an agar plate with bacteria isolated from a patient's infection.
However, the method has been developed, refined and standardized by many scientists and scientific organizations over the years including George F. Reddish, Norman Heatley, James G. Vincent,[8] Alfred W. Bauer, William M.M.
[8] A pure bacterial culture is suspended in saline, its turbidity is standardized, and it is swabbed uniformly across an agar plate.
In general, larger zones of inhibition correlate with lower minimum inhibitory concentrations (MICs) of antibiotic or extract for that bacterial strain.
Bacterial inoculum is prepared by diluting a broth culture to match a 0.5 McFarland turbidity standard, which is equivalent to approximately 150 million cells per mL.
In the case of Gram negative bacteria, excess liquid is removed from the swab by gently pressing or rotating it against the inside of the tube.
[6][11] Disks containing increasing antibiotic concentrations are placed on a seeded bacterial lawn on the agar surface and plates are incubated.
These are plotted as linear dimensions or squares of distances as a function of the natural logarithm of antibiotic concentration in the disks.