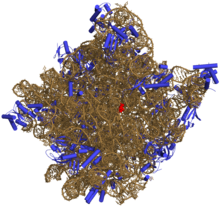
Prokaryotic large ribosomal subunit
X-ray crystallography has yielded electron density maps allowing the structure of the 50S in Haloarcula marismortui (archaeon) to be determined to 2.4Å resolution[1]and of the 50S in the Deinococcus radiodurans (bacterium) to 3.3Å.
[6] A cryoEM reconstruction of the native 50S subunit of the extremely halophilic Archaean Halococcus morrhuae (classified under Euryarchaeota; Stenosarchaea group) is available.
[8][9] 50S includes the activity that catalyzes peptide bond formation (peptidyl transfer reaction), prevents premature polypeptide hydrolysis, provides a binding site for the G-protein factors (assists initiation, elongation, and termination), and helps protein folding after synthesis.
CCA sequence at its acceptor stem is bound to the A site, C74 of the tRNA stacking with U2590 (2555) induces a conformational change in the ribosome, resulting in movement of U2541 (2506), U2620 (2585) through G2618 (2583).
The N3 (nitrogen) of A2486 (2451) is closest to the peptide bond being synthesized and may function as a general base to facilitate the nucleophilic attack by the amino group of the aminoacyl-tRNA (in the A site).
This buried phosphate can stabilize the normally rare imino tautomers of both bases, resulting in an increase in the negative charge density on N3.
After initiation, elongation, and termination, there is a fourth step of the disassembly of the post-termination complex of ribosome, mRNA, and tRNA, which is a prerequisite for the next round of protein synthesis.